Crash course on R markdown
Overview
The research world (and eventually the teaching world as well) is moving very fast towards requiring reproducible research1–6. “The ultimate product of academic research is the paper along with the laboratory notebooks and full computational environment used to produce the results in the paper such as the code, data, etc. that can be used to reproduce the results and create new work based on the research” (Wikipedia).
The consequences are that we should change our habits and certainly start preparing all our manuscripts, presentations, homework, etc., in a format that is clean and reproducible, i.e., if you give anyone the code, this person can reproduce the document exactly. This document seeks to make this transition somewhat easier using R Markdown.
What is the Markdown language?
I borrow much of the very good text from Dr Shalizi from Carnegie Mellon University about Mark-up and Markdown.
Many word processing programs, like Microsoft Word, employ the “what you see is what you get” (WYSIWYG) principle: you want some words to be printed in italics? You select/highlight the words or phrase, do ctrl-I or click Italic, and they’re in italics right there on the screen. You want some words to be in a bigger, different font, so you just select the font, and so on. This works well enough for the vast majority of things that are still currently done but is not a reproducible basis for a system of text formatting, because it depends on a particular program (a) knowing what you mean and (b) implementing it well.
For several decades, really serious systems for writing have been based on a very different principle, that of marking up text. The essential idea in a mark-up language is that it consists of ordinary text, plus signs which indicate how to change the formatting or meaning of the text. Some mark-up languages, like HTML (Hyper-Text Markup Language) use very obtrusive markup; others, like the language called Markdown, are more subtle.
Obviously, there has been a play-on-word between Mark-up and Markdown…!
Much of the content of the paragraphs above looks like this in HTML:
<div id="what-is-the-markdown-language" class="section level2">
<h2>What is the Markdown language?</h2>
<p>I borrow ... from <a href="http://www.stat.cmu.edu/~cshalizi/rmarkdown/">Dr Shalizi</a> from ...</p>
<p>Many ...that of <em>marking up</em> text. The essential idea in a <strong>mark-up language</strong> is that it consists of ordinary text, <em>plus</em> signs which indicate how to change the formatting or meaning of the text. Some mark-up languages, like HTML (Hyper-Text Markup Language) use very obtrusive markup; others, like the language called <strong>Markdown</strong>, are more subtle.</p>
</div>and like this in Markdown:
## What is the Markdown language?
I borrow ... from [Dr Shalizi](http://www.stat.cmu.edu/~cshalizi/rmarkdown/) from ...
Many ... that of _marking up_ text. The essential idea in a **mark-up language** is that it consists of ordinary text, _plus_ signs which indicate how to change the formatting or meaning of the text. Some mark-up languages, like HTML (Hyper-Text Markup Language) use very obtrusive markup; others, like the language called **Markdown**, are more subtle.Every mark-up language needs to be rendered somehow into a format which actually includes the fancy formatting, images, mathematics, computer code, etc., etc., specified by the mark-up. For HTML, the rendering program is called a “web browser”. Most computers which know how to work with Markdown at all know how to render it as HTML (which you can then view in a browser), PDF (which you can then view in Acrobat or the like), or Word (which you can then view in Microsoft Word).
The advantages of mark-up languages are many: they tend to be more portable across machines, less beholden to particular software companies, and more stable over time than WYSIWYG word processing programs. R Markdown is, in particular, both “free as in beer” (you will never pay a dollar for software to use it) and “free as in speech” (the specification is completely open to all to inspect). The sheer stability of mark-up languages makes them superior for scientific documents.
Markdown is thus a simple mark-up language with the goal of enabling people “to write using an easy-to-read, easy-to-write plain text format, and optionally convert it to structurally valid XHTML (or HTML)” (wikipedia). OK, so imagine that this is a writing language that is very close to plain text, where the most useful alteration of text or mark-up (headings, bold, italic, web references, etc.) are simpler than the html codes. I encourage you to read the wikipedia definition of Markdown to have more details of why and how it came about.
The ‘R twist’ of Markdown: R Markdown
Now the beauty of Markdown is that some very smart people have thought that one could in fact embed R codes, within the text, such that in the end, a document could contain all the text and code to create figures and tables. Other, just as smart people, have thought that one could use all the beauty of the LaTeX, particularly for editing equations. They have also added the possibility to add citations, bibliography. All this to say that they have added all the features that students/researchers need and have dreamt about for a while!
So the ‘R twist’ to Markdown is called ‘R Markdown’. One of the hidden beauties is that R Mardown uses Pandoc, which is a universal document converter, to render the text. The result is that you and I will now write in R Markdown and will have the possibility of publishing what we have written in many output formats, which include among many others HTML, pdf, and Word. This is one of the major advantages compared to LaTeX, which can only produce pdf documents.
R Markdown in R Studio
I have had a very low opinion of R studio until very recently because I never liked the text editor, I always thought (still do in fact…!) that on a normal screen, things were just too cramped. But, I must say that I have come to change my mind, principally thanks to the possibilities that R Markdown offers, and because R Mardown is hosted in R studio.
The goal of this document is thus to introduce you to R Markdown and the things you need to get to know quickly for authoring HTML, PDF, MS Word documents, but also many other interesting outputs. All the details on using R Markdown can be found at http://rmarkdown.rstudio.com. I encourage you to browse the gallery and then spend time to get familiarized on what can be done with it on the step by step tutorial.
I still think that there is nothing like playing with examples to make sense of things. I assume that you have now browsed through the guide reference and you have installed the rmarkdown and knitr packages in RStudio. If not, in the console, type install.packages("rmarkdown","knitr"). First, you want to open a Rmd document. Go to File -> New File -> R Markdown... and click OK with the default variables. Click the Knit button just above the window where the document just appeared. Another ‘Choose Encoding’ pop-up window will appear, click OK with the default values. The system will ask you where you want to save this document and under what name. Then, an html preview pop-up window should appear where there is headings, text, hyperlinks, results from some R commands and a simple plot. OK, time to look at the .Rmd file content.
Components of R Markdown
There are three components to an R Mardown document (.Rmd):
- YAML header
- Text
- Code chunks
After you are done reading this document, you will want to be able to find a reference summary of R markdown here.
- YAML header: The header of the document starts with a YAML which acronym started as “Yet Another Markup Language” and now apparently stands for “YAML Ain’t Markup Language”… The YAML Header specifies how the document will be formatted.
- Text: most of your writing will be done using plain text, very much like you have been doing with Word, with mark-up alterations. Many details on mark-up are in the section devoted to text below. Spell check can be done using
- A spell check button to the right of the save button (with “ABC” and a check mark).
- Edit > Check Spelling…
- The F7 key
- Code Chunks: This is where you will add the actual
Rcode that will create tables and figures. The details are in the section devoted to Code Chunks below
YAML header
The header of the document starts with a YAML which acronym started as “Yet Another Markup Language” and now apparently stands for “YAML Ain’t Markup Language”… The YAML Header specifies how the document will be formatted. For example, in this case, we have specified that the output will be an HTML document. Yours should look very close to something like this:
---
title: "Untitled"
author: "François Birgand"
date: "12/10/2017"
output: html_document
---The title will appear as your title document. It is unrelated to the name of your file. If you are writing a paper, this will be the title of your paper.
The author will appear when you render or ‘knit’ your document under the title and introduced by a “by”… The date corresponds to the initial date of creation. If you want to use the date at the time you knit the document, you may consider putting an inline code (details below) such as `r format(Sys.Date() `%d %B %Y`)`.
Output document in YAML
You can change the output: html_document, to output: github_document, to output: word_document, to output: pdf_document. If you are having trouble with the pdf_document output, this means that the LaTeX1 engine is not installed on your computer. For this I am borrowing instructions from here where they suggest these links:
- For Windows: Go to https://miktex.org/howto/install-miktex and follow directions to install MikTex.
- For Mac: Go to https://tug.org/mactex/mactex-download.html and follow directions to install MacTex.
In the YAML, one puts some broad instructions on how Pandoc needs to interpret the document, one big information being the output. You can see right away that with just one document, you can create at least 4 different documents. when you typed output: github_document, there was an html preview that popped-up, but in the background, a *.md file was created. This *.md* file is interpreted by GitHub to render the document within the GitHub environment. If you try to click on a *.html file that is hosted on GitHub, only code will appear, and not the rendered html document.
Table Of Content
R markdown provides the possibility of Table Of Content or toc that you put as a variable in the YAML part of the R Markdown. The toc will appear at the head of the document and will give you links to all the heading levels you have created. This is very useful to quickly give you an outline of your document and allows you to get directly to the section of your choice. The YAML of this document is
---
title: "Crash course on R Markdown"
author: "François Birgand"
date: "09/10/2017"
output:
html_document:
toc: true
toc_float: true
bibliography: RMD_course_bibtex.bibtex
link-citations: true
csl: journal-of-hydrology.csl
---Notice that the html_document has now been put on another line for organization purposes, but also note that a : has been added at the end of html_document:. This tells Pandoc that among the html_document, a toc should be added.
Bibliography
I have created and maintained for over 15 years a database written in Microsoft Access of around 2000 articles, most of them photocopied and typed in the database by hand during and after my Ph.D. (yes, I should have a medal!…) but I have to admit that this system is past its time… I have now switched to Paperpile. There are many other referencing software out there now (a list and comparison can be found here). The beauty of them is that they will all export in recognized formats and this is what we care about here. So no more copy/paste of references from some pdf. We are going into reproducible research. This means that all the articles you want to work with will be entered in the reference database of your choice. Make sure you start one ASAP!
In the YAML header, you can now see bibliography: RMD_course_bibtex.bibtex. This means that there is a file named RMD_course_bibtex.bibtex that is located in the same directory as the .Rmd file, which when we ‘knit’, Pandoc will look for. You can access this .bibtex file in the GitHub directory. There is actually a possibility to add within the .Rmd document itself, the text needed for Pandoc to read the references, but unless you only have very few references, this can become very messy quickly. See the RStudio Bibliographies documentation for more details.
In short, in this file, there is a list of articles which information is coded according to the format chosen. By the way, a lot of the details for the bibliography are available on the RStudio Bibliographies documentation. R Markdown will recognize up to 10 different formats. The one Paperpile exports is .bibtex, so this is the one I have have added in the YAML header. The filename extension will determine the rendering process, so make sure you have the right extension as well.
So, in the .bibtex file I have, the first article appears as
@ARTICLE{Kuhne2009-tn,
title = "Improving the Traditional Information Management in Natural
Sciences",
author = "K{\"u}hne, Martin and Liehr, Andreas W",
journal = "Data Sci. J.",
volume = 8,
pages = "18--26",
year = 2009,
issn = "1683-1470, 0308-9541",
doi = "10.2481/dsj.8.18"
}The first item after “ARTICLE{” is the unique identifier for the article. The identifier of this article is Kuhne2009-tn. When one wants to cite this article, one will say something like “reproducible research has been suggested to become the norm2”. And you would code like
"reproducible research has been suggested to become the norm [@Kuhne2009-tn]"If you want to say that “Kühne et al.2 have shown that etc.”, you would add a “dash” just before the “@” and code it as such:
Kühne et al. [-@Kuhne2009-tn] have shown that etc."If you want to cite several references, you would add a semicolon between the two such as in:
"reproducible research has been suggested to become the norm [@Kuhne2009-tn; @Buckheit1995-ls]"Notice that among all the fields from the example above, there are doi and issn. doi stands for “Digital Object Identifier”. issn stands for International Standard Serial Number (ISSN), which is an eight-digit serial number used to uniquely identify a serial publication. The doi value in this example is 10.2481/dsj.8.18, which is unique in the world. These doi are applied to articles but not only. They are also applied to data. Eventually, all data that will be used in an article will have to have its own doi and all the codes that are used to analyze the data will refer to this unique doi. This is not quite implemented yet (in 2017), but will likely be by 2022. doi or url (not added here; stands for Uniform Resource Locator. Quite a mouthful, really, for what it is: a web address) are not necessarily exported from your reference software by default. Make sure you add this possibility, as doi is almost routinely added in the reference list in most journals.
The reference list of the citations will appear right after you place
# Referencesat the bottom of your Rmd docucment. When rendering your document, the list will appear automatically afterwards, and if you have in text notes, these will appear underneath.
Link to citations
One very nice feature is to create hyperlinks from the in-text citations to the citations in the reference section. For this, just add link-citations: true in the YAML header.
CSL and styling of citations
CSL stands for Citation Style Language. The CSL line command is an option for the citation styles to be used in your document. You can comment it out by adding a “#” in front of it and the default .CSL file will apply without you noticing it. Each journal has its own way of handling how an article/reference should be cited in the text, and in the reference section, and there are hundreds of different styles out there… You can read lots of details on this CSL primer about how all this works.
While I was writing this tutorial, I did not specify at first the citation style. And I kept getting, using the same example as before, “(Kühne and Liehr 2009)”, although I wanted “(Kühne and Liehr, 2009)”, i.e., with a comma between the authors and the date, because this is the way I always did it and I think it is a lot better this way. Then I started thinking about the journals for which the inline citations are numbers, sometimes in brackets, sometimes without brackets… what a nightmare…! Actually it is extremely simple: all this is done automatically when you specify the CSL corresponding to the journal style following which you wish to write.
For this, you can pick at https://github.com/citation-style-language/styles the *.CSL file you are looking for (actually there are too many of them and they are not all displayed). The fastest way is to Google “raw icon on GitHub, and copy all the file in a text editor. Warning! You need to make sure that your text editor does not add any weird formatting or add an extension at the end of the file. I had that problem and I could not see the extension on my computer, although I had the option to display so. Save the file in the same directory as your .Rmd.
So, to go back with my struggling to style the inline citation and the “missing comma”, it turns out that the default CSL file uses a “Chicago author-date format” (I am not sure what this means exactly), which in text styling is “(author date)”, without the comma…! If you use “journal-of-hydrology.CSL”, you will see that all of a sudden after you knit, the commas just appeared…! Eureka! If you use “nature.CSL” (because we all want to be ready when we publish in Nature!), you will see that there are no more in-line citations, just numbers in superscript, hyphenated when needed, and the references are all in order, with the journal names in italics, the journal volume in bold, the year in parenthesis at the end, etc.! Is not that wonderful? Without your doing anything, other than adding the citation properly in the text following the guidelines above.
You can, if you want and have a lot of time to waste, take existing CSL files and modify them to have your own custom citation style. Make sure you take an “independent” CSL file and modify it. Most of them are dependent upon a “source” or independent one, and the code in the CSL file is just saying how the dependent file should slightly alter the independent one. Good instructions on how to do this can be found on the excellent CSL primer.
Text
The markdown syntax is very simple and this is one of the reasons why it is so attractive. You can find all the syntax with this R Markdown reference document, but I thought I would go through them as I have learned several tricks I could not easily find in the document.
Paragraph Breaks and Forced Line Breaks
To insert a break between paragraphs, include a single completely blank line.
To force a line break, put two blank spaces at the end of a line.
To insert a break between paragraphs, include a single completely blank line.
To force a line break, put _two_ blank spaces at the end of a line.Headers
The character # at the beginning of a line means that the rest of the line is interpreted as a section header. The number of #s at the beginning of the line indicates the level of the section (1,2,3, etc.). For instance, Components of R Markdown above is preceded by a single #, because it is of level 1 but Headers at the start of this paragraph is preceded by ### because it is of level 3. Up to six levels are understood by Markdown. Do not interrupt these headers by line-breaks. Make sure that in your .Rmd file, you leave a blank line before a header, otherwise, pandoc will not render it as a header.
Italics and bold
*italics* and _italics_renders italics and italics
**bold** and __bold__renders bold and bold
Supbscripts and superscripts
To write sub- and superscripts, like in NO3- or PO43- write as
NO~3~^-^ or PO~4~^3-^PO43- does not look as neat as \(PO_4^{3-}\)
PO~4~^3-^ does not look as neat as $PO_4^{3-}$but looks more seamless in a normal text because the former does not appear as an equation while the other does.
Lists
* unordered list
* item 2
+ sub-item 1 #4 <spaces> before +
+ sub-item 2 #4 <spaces> before +- unordered list
- item 2
- sub-item 1
- sub-item 2
1. ordered list
2. item 2
+ sub-item 1 #4 <spaces> before +
+ sub-item 2 #4 <spaces> before +- ordered list
- item 2
- sub-item 1
- sub-item 2
In the ordered list, there is a subtlety unrevealed in the Rmardown documentation, which is that the numbering always increases, and that only the number value of the first item matters. So, one cannot have a list in a decreasing order (which is too bad because when one makes a list of his/her publications, it is nice to have a decreasing order…), and the only number that matters is the first one. So this code
5. ordered list
7. item 2
2. item 2
# blank line for list to take into effect
b. sub-item 1 #4 <spaces> before b
a. sub-item 2
renders this:
- ordered list
- item 2
item 2
- sub-item 1
- sub-item 2
- sub-item 1
Quotations
R Markdown is, in particular, both “free as in beer” (you will never pay a dollar for software to use it) and “free as in speech” (the specification is completely open to all to inspect).
Is a quotation from Dr Shalizi which code starts with a >
> R Markdown is, in particular, both "free as in beer" etc.Computer type
This is to differentiate regular text from code text so that both can be easily differentiated: R vs R.
This is to differentiate regular text from `code text` so that both can be easily differentiated: `R` vs R.An entire paragraph of code which is rendered as a “code box” in html (but not in pdf) starts with three “back-ticks”, and end the same
Symbols and Special characters
The principal keys, like the alphabet, are understood univocally across platforms such as Windows, Mac OS, or Linux. However, there are special characters such °, ² or µ that have different embedded codes across the different platforms. For example, if you and your co-worker work on the same document and one works using Windows and the other uses Mac, the actual symbol in the code text may not show the same one from one to the other plateform.
For example, if I write in an R markdown document “10 m²” and I have added the ² symbol by typing on a PC “Alt+0178”, as this corresponds to the ascii code for ² in Windows, the same document open on a Mac will render “10 m?”, because it cannot interpret the embedded code for ²…
Several consequences:
- NEVER use special characters in variable names in an
Rcode, - in an R markdown document, use the HTML Number or HTML name in the text
HTML Number and Names can be found on this ascii page
So to type T°C, m² or µm you type like this
So to type T°C, m² or µm you type like thisDo not forget to add the “;” !!!
Absolute Links
a link within the text as a [link](www.rstudio.com)a link within the text as a link. Absolute links to images, see Images below.
Links within the document
Linking to a heading
If you want to provide a link (other than the Table Of Content at the beginning of the document) to an existing heading like The ‘R twist’ of Markdown: R Markdown, you type it as below
If you want to provide a link back to an existing heading like [The 'R twist' of Mardown: R Markdown](#the-r-twist-of-markdown-r-markdown), you type it as belowIt provides a link using the regular [text](*address of text*) syntax for a link. Now, this time the *address of text* is not an absolute address as shown in the link example, but rather a relative one which has a format like (#...). Mardown took the name of the heading The 'R twist' of Markdown: R Markdown and created for each heading what it referred to as a <div> in HTML and it gave it an identity or id, and for this example here, it gave in HTML code <div id="the-r-twist-of-markdown-r-markdown" class="section level1">. It is important to notice that the original text of the heading "The 'R twist' of Markdown: R Markdown" is transformed into "the-r-twist-of-markdown-r-markdown" where all signs have been removed, spaces replaced by a "-", and all characters are in lower case.
Linking to anywhere in the document
There is, to my knowledge, no native way of doing this in Markdown. One must borrow some HTML code and it is very simple. You must add an anchor link in your document. where you want the page to line up. So in the link example example above point to the absolute link example, the code is like that.
So in the [link example](#link_ref) example above point to the absolute link example, the code is like that.I thus added in the line just above where I want the link this HTML code where <a> stands for anchor.
<a id="link_ref"></a>Footnotes
You can add footnotes in the text by adding [^footnote] right next to the word concerned and add the [^footnote]: with the text next to it JUST BEFORE the # References section.
Images
To add images in a document, add a ! in front of the same syntax as for links such as:
to render:

You can modify the width of the image by adding e.g., {width = 50%} to have the image take 50% of the width of the page
{width=50%}to render:

To align the image, there is no native way of doing it in Markdown, so you can use a little bit of HTML code like <center> ... </center> such as below. Apparently, it might not work that great in html5 though…
<center>{width=25%}</center>to render:

You can add a caption by just adding the “Using a caption this time” in the brackets.

Using a caption this time
{width=33%}Now, you can add an image in the text like this  and it will work just fine.
and it will work just fine.
Now, you can add an image in the text like this {width=33%} and it will work just fine. However, you can see that the caption was not added. So for the caption to be added, there needs to be a return before and after the image code. If you include the image like this

The caption does not appear.
However, you can see that the caption was not added. So for the caption to be added, there needs to be a return before and after the image code. If you include the image like this
{width=33%}
The caption does not appear.But if you leave a blank line before and after like this

Using a caption this time
The caption will appear.
But if you leave a blank line before and after like this
{width=33%}
The caption will appear.Now if you want to center both figure and caption, you need to add <center> a blank line, the image code, a blank line, then </center>, and it will appear like this:

Using a caption this time
<center>
{width=33%}
</center>And you can see that adding *__ __* aound the caption text put the caption in bold and italic as well.
To have an image linked to a particular website, you can use this code:
[](https://www.bae.ncsu.edu/wp-content/themes/riddick/dist/images/ncstate-logos/ncstate-brick-2x2-red.png)If you do that, however, the caption will not show any more, even though you have the proper number of lines before and after. If you want a caption below with a link, you will have to add this underneath. So if you want both the image and the caption to have clickable links, and centered like this
you would write:
<center>
[{width=33%}](https://www.bae.ncsu.edu/wp-content/themes/riddick/dist/images/ncstate-logos/ncstate-brick-2x2-red.png)
[This is the corresponding clickable caption I want](https://www.bae.ncsu.edu/wp-content/themes/riddick/dist/images/ncstate-logos/ncstate-brick-2x2-red.png)
</center>Editing equations in R Markdown
We do not write nearly as much math in our field than our colleagues in statistics or math. But we do enough to have a bit of a tutorial. I have borrowed this entire section from the excellent tutorial from Dr Shalizi again. R Markdown gives you the syntax to render complex mathematical formulas and derivations, and have them displayed very nicely. Like code, the math can either be inline or set off (displays).
Inline equations
Inline math is marked off witha pair of dollar signs ($), as \(\pi r^2\) or \(e^{i\pi}\).
Inline math is marked off witha pair of dollar
signs (`$`), as $\pi r^2$ or $e^{i\pi}$.Set off equations
Mathematical displays are marked off with \[ and \], as in \[
e^{i \pi} = -1
\]
Mathematical displays are marked off with `\[` and `\]`, as in
\[
e^{i \pi} = -1
\]Once your text has entered math mode, R Markdown turns over the job of converting your text into math to a different program, called LaTeX. This is the most common system for typesetting mathematical documents throughout the sciences, and has been for decades. It is extremely powerful, stable, available on basically every computer, and completely free. It is also, in its full power, pretty complicated. Fortunately, the most useful bits, for our purposes, are actually rather straightforward.
Elements of Math Mode
- Most letters will be rendered in italics (compare: a vs.
avs. \(a\); only the last is in math mode, i.e.,$a$). The spacing between letters also follows the conventions for math, so don’t treat it as just another way of getting italics. (Compare speed, in simple italics, with \(speed\), in math mode, i.e.,$speed$) - Greek letters can be accessed with the slash in front of their names, as
\alphafor \(\alpha\). Making the first letter upper case gives the upper-case letter, as in\Gammafor \(\Gamma\) vs.\gammafor \(\gamma\). (Upper-case alpha and beta are the same as Roman A and B, so no special commands for them.) - There are other “slashed” (or “escaped”) commands for other mathematical symbols:
\timesfor \(\times\)\cdotfor \(\cdot\)\leqand\geqfor \(\leq\) and \(\geq\)\subsetand\subseteqfor \(\subset\) and \(\subseteq\)\leftarrow,\rightarrow,\Leftarrow,\Rightarrow,\rightleftharpoonsfor \(\leftarrow\), \(\rightarrow\), \(\Leftarrow\), \(\Rightarrow\), \(\rightleftharpoons\)\approx,\sim,\equivfor \(\approx\), \(\sim\), \(\equiv\)- See, e.g., http://web.ift.uib.no/Teori/KURS/WRK/TeX/symALL.html for a fuller listing of available symbols. (http://tug.ctan.org/info/symbols/comprehensive/symbols-a4.pdf lists all symbols available in
LaTeX, including many non-mathematical special chracters)
- Subscripts go after an underscore character,
_, and superscripts go after a caret,^, as\beta_1for \(\beta_1\) ora^2for \(a^2\). - Curly braces are used to create groupings that should be kept together, e.g.,
a_{ij}for \(a_{ij}\) (vs.a_ijfor \(a_ij\)). - If you need something set in ordinary (Roman) type within math mode, use
\mathrm, ast_{\mathrm{in}}^2for \(t_{\mathrm{in}}^2\). - If you’d like something set in an outline font (“blackboard bold”), use
\mathbb, as\mathbb{R}for \(\mathbb{R}\). For bold face, use
\mathbf, as(\mathbf{x}^T\mathbf{x})^{-1}\mathbf{x}^T\mathbf{y}for \[ (\mathbf{x}^T\mathbf{x})^{-1}\mathbf{x}^T\mathbf{y} \]
- Accents on characters work rather like changes of font:
\vec{a}produces \(\vec{a}\),\hat{a}produces \(\hat{a}\). Some accents, particularly hats, work better if they space out, as with\widehat{\mathrm{Var}}producing \(\widehat{\mathrm{Var}}\). - Function names are typically written in romans, and spaced differently: thus \(\log{x}\), not \(log x\).
LaTeX, and thereforeR Markdown, knows about a lot of such functions, and their names all begin with\. For instance:\log,\sin,\cos,\exp,\min, etc. Follow these function names with the argument in curly braces; this helpsLaTeXfigure out what exactly the argument is, and keep it grouped together with the function name when it’s laying out the text. Thus\log{(x+1)}is better than\log (x+1). Fractions can be created with
\frac, like so:
produces \[ \frac{a+b}{b} = 1 + \frac{a}{b} \]\frac{a+b}{b} = 1 + \frac{a}{b}Sums can be written like so:
will produce \[ \sum_{i=1}^{n}{x_i^2} \] The lower and upper limits of summation after the\sum_{i=1}^{n}{x_i^2}\sumare both optional. Products and integrals work similarly, only with\prodand\int: \[ n! = \prod_{i=1}^{n}{i} \] \[ \log{b} - \log{a} = \int_{x=a}^{x=b}{\frac{1}{x} dx} \]\sum,\prodand\intall automatically adjust to the size of the expression being summed, producted or integrated.“Delimiters”, like parentheses or braces, can automatically re-size to match what they’re surrounding. To do this, you need to use
\leftand\right, as
renders as \[ \left( \sum_{i=1}^{n}{i} \right)^2 = \left( \frac{n(n-1)}{2}\right)^2 = \frac{n^2(n-1)^2}{4} \]\left( \sum_{i=1}^{n}{i} \right)^2 = \left( \frac{n(n-1)}{2}\right)^2 = \frac{n^2(n-1)^2}{4}- To use curly braces as delimiters, precede them with slashes, as
\{and\}for \(\{\) and \(\}\). Multiple equations, with their equals signs lined up, can be created using
eqnarray, as follows.\[ \begin{eqnarray} X & \sim & \mathrm{N}(0,1)\\ Y & \sim & \chi^2_{n-p}\\ R & \equiv & X/Y \sim t_{n-p} \end{eqnarray} \]
\[ \begin{eqnarray} X & \sim & \mathrm{N}(0,1)\\ Y & \sim & \chi^2_{n-p}\\ R & \equiv & X/Y \sim t_{n-p} \end{eqnarray} \]
Notice that & surrounds what goes in the middle on each line, and each line (except the last) is terminated with \\.
Code in “Chunks” and within the text
The real beauty of R Markdown is the ability of adding R codes that can be rendered or interpreted along with the mark-up text. R code can be inserted as “chunks” and “in-line”.
Inline code
You can embed some r code within the text by using `r signif(pi,5)` to give 5 significant digits to the number $\pi$ You can embed some r code within the text by using 3.1416 to give 5 significant digits to the number \(\pi\).
Code chunks
A code chunk is simply an off-set piece of code by itself. It is preceded by ```{r} on a line by itself, and ended by a line which just says ```. The code itself goes in between.
```{r cars}
summary(cars)
```generates these results:
## speed dist
## Min. : 4.0 Min. : 2.00
## 1st Qu.:12.0 1st Qu.: 26.00
## Median :15.0 Median : 36.00
## Mean :15.4 Mean : 42.98
## 3rd Qu.:19.0 3rd Qu.: 56.00
## Max. :25.0 Max. :120.00First, in the syntax, the word “cars” after the “{r” is the name of the chunk (see below). To let the renderer program know that this is now a R code, the letter “r” needs to follow the “{”.
You can also embed plots, for example:
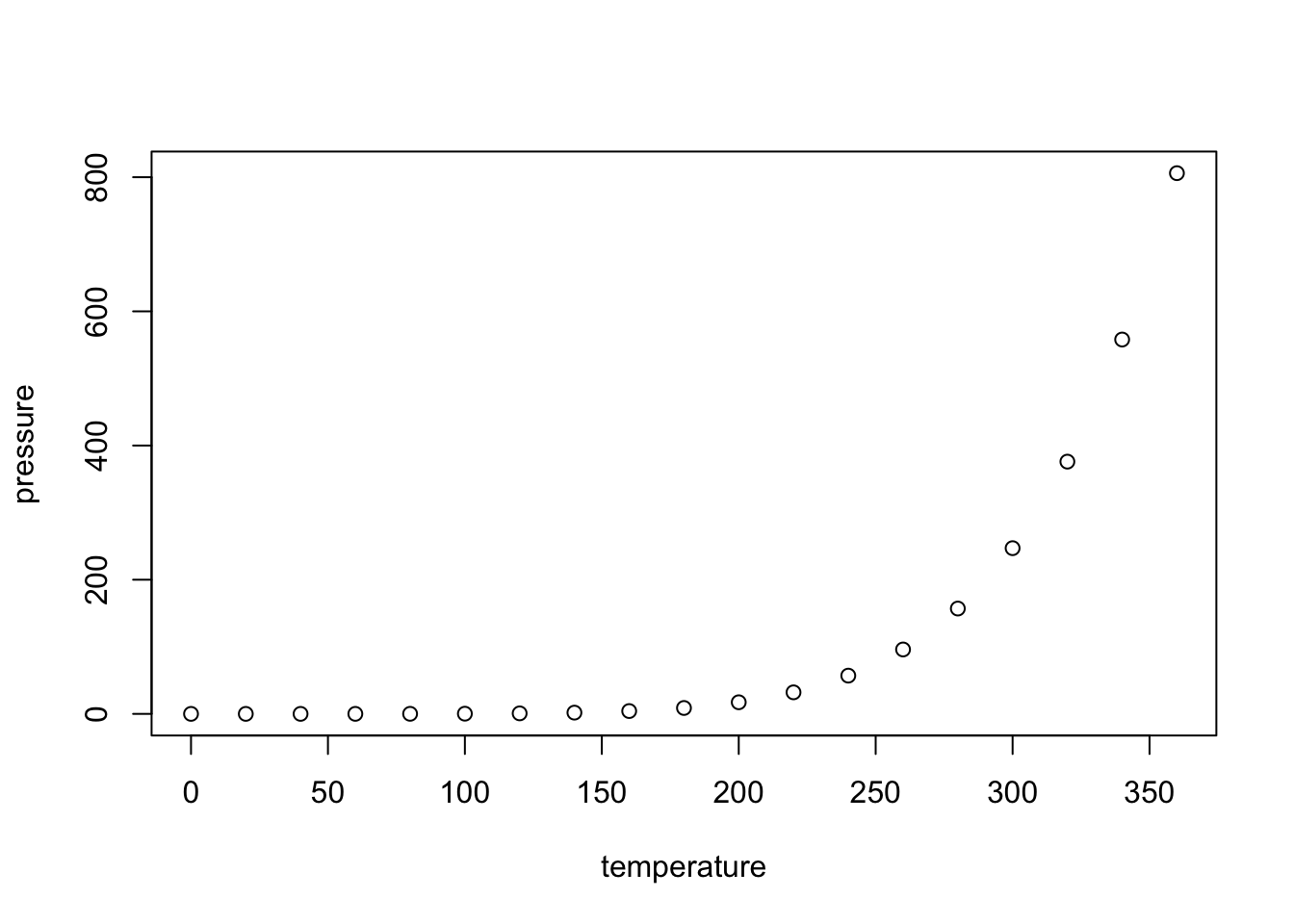
Markdown tables and the “kable” function
coming soon…
Chunk results options
Code chunks (but not inline code) can take a lot of options which modify how they are run, and how they appear in the document. These options go after the initial r and before the closing } that announces the start of a code chunk. One of the most common options turns off printing out the code, but leaves the results alone: ```{r, echo=FALSE}
Another runs the code, but includes neither the text of the code nor its output. ```{r, include=FALSE} This might seem pointless, but it can be useful for code chunks which do set-up like loading data files, or initial model estimates, etc.
Another option prints the code in the document, but does not run it: ```{r, eval=FALSE} This is useful if you want to talk about the (nicely formatted) code.
Another option on the results of the code is that it generate all results “as-is”, which is very nice when your code generates mark-up text to be rendered by Pandoc.
```{r, results="asis"}
By default, the results of a chunk with have ## as a prefix. You can remove this by putting
```{r, comment=FALSE}
Sometimes, running of the code will generate warnings and messages. These can be turned off in the output by using
```{r, warning=FALSE, message = FALSE}
Naming Chunks
You can give chunks names immediately after their opening, like ```{r, clevername}. This name is then used for the images (or other files) that are generated when the document is rendered.
Adjusting figure sizes and alignments
These details are discussed in the accompanying written article on instantaneous vs. interval-average flow data.
“Caching” Code Chunks (Re-Running Only When Changed)
By default, R Markdown will re-run all of your code every time you render your document. If some of your code is slow, this can add up to a lot of time. You can, however, ask R Markdown to keep track of whether a chunk of code has changed, and only re-run it if it has. This is called caching the chunk.
```{r, cache=TRUE}
summary(cars)
```## speed dist
## Min. : 4.0 Min. : 2.00
## 1st Qu.:12.0 1st Qu.: 26.00
## Median :15.0 Median : 36.00
## Mean :15.4 Mean : 42.98
## 3rd Qu.:19.0 3rd Qu.: 56.00
## Max. :25.0 Max. :120.00One issue is that a chunk of code which hasn’t changed itself might call on results of earlier, modified chunks, and then we would want to re-run the downstream chunks. There are options for manually telling R Markdown “this chunk depends on this earlier chunk”, but it’s generally easier to let it take care of that, by setting the autodep=TRUE option.
- If you load a package with the
library()command, R Markdown isn’t smart enough to check whether the package has changed (or indeed been installed, if you were missing it). So that won’t trigger an automatic re-running of a cached code chunk. - To manually force re-running all code chunks, the easiest thing to do is to delete the directory R Markdown will create (named something like filename
_cache) which it uses to store the state of all code chunks.
Setting Defaults for All Chunks
You can tell R to set some defaults to apply to all chunks where you don’t specifically over-ride them. Here are the ones I generally use:
```{r, eval=FALSE}
# Need the knitr package to set chunk options
library(knitr)
# Set knitr options for knitting code into the report:
# - Don't print out code (echo)
# - Save results so that code blocks aren't re-run unless code changes (cache),
# _or_ a relevant earlier code block changed (autodep), but don't re-run if the
# only thing that changed was the comments (cache.comments)
# - show the error messages (message)
# - Don't clutter R output with warnings (warning)
# This _will_ leave error messages showing up in the knitted report
opts_chunk$set(echo=FALSE,
cache=TRUE, autodep=TRUE, cache.comments=FALSE,
message=TRUE, warning=FALSE)
```This sets some additional options beyond the ones I’ve discussed, like not re-running a chunk if only the comments have changed (cache.comments = FALSE), and leaving out messages and warnings. (I’d only recommend suppressing warnings once you’re sure your code is in good shape.) I would typically give this set-up chunk itself the option include=FALSE.
You can over-ride these defaults by setting options for individual chunks.
More Chunk options
See [http://yihui.name/knitr/options/] for a complete listing of possible chunk options.
Additional Subtleties
Creating anchors and links within a page
Now, further down in the document, you might be interested in linking back up if you want to make a link to a particular heading or a particular place in the document that is not a heading.
Challenges with the “back-ticks”
As I was writing this document, I wanted some of the code to appear as I wrote it, in the “code frames”. It turns out that to illustrate that to run a code in line the text, one needs to use `r signif(pi,5)` , well it is not simple because the ` is interpreted as containing some code! Therefore, it is not interpreted as text not intended to be rendered…!! So it is a bit of a tour de force to make it appear and being uninterpreted. It turns that in R, if you type
'\x60'it will return the “back-tick” symbol `. But to display it I have to type in the text
== =r '\x60'= ==where for illustration purposes, I have replaced the “back-ticks”, by the “=” signs. For another example to display `r signif(pi,5)` in the text, I actually coded
== =r '\x60r signif(pi,5)\x60'= ==where again I have replaced the “back-ticks” with “=” signs.
References
1. Buckheit, J. B. & Donoho, D. L. Wavelab and reproducible research. (Stanford University, Standford CA 94305, USA, 1995).
2. Kühne, M. & Liehr, A. W. Improving the traditional information management in natural sciences. Data Sci. J. 8, 18–26 (2009).
3. the Yale Law School Roundtable on Data and Code Sharing. Reproducible research. Computing in Science Engineering 12, 8–13 (2010).
4. Wicherts, J. M. & Bakker, M. Publish (your data) or (let the data) perish! Why not publish your data too? Intelligence 40, 73–76 (2012).
5. Pasquier, T. et al. If these data could talk. Sci Data 4, 170114 (2017).
6. Peng, R. D. Reproducible research in computational science. Science 334, 1226–1227 (2011).
In the 1970s, the great computer scientist Donald Knuth wrote a mark-up language, and a rendering program for that language, called
TeX(pronounced “tech”), for writing complex mathematical documents. In the 1980s, the computer scientist Leslie Lamport extendedTeXin ways that made it rather more user-friendly, and called the resultLaTeX(pronounced “la-tech”)↩